Document Type : Short Communication
Authors
1 Department of Clinical Sciences, School of Veterinary Medicine, Shiraz University, Shiraz, Iran
2 Department of Pathobiology, School of Veterinary Medicine, Shiraz University, Shiraz, Iran
Keywords
Introduction
Echinococcosis infection is a cosmopolitan zoonosis caused by the adult or larval stages of cestodes belonging to the genus Echinococcus (family: Taeniidae). The disease is one of the more prevalent infections in Iran especially in rural areas, where offal from abattoirs is improperly disposed, or where slaughtering is done on farms.1 The parasite has an indirect life cycle. Adult worms occur in dogs and other canids as definitive hosts and many herbivorous and omnivorous species, including sheep, cattle, camels, pigs and humans as intermediate hosts.2 Hydatidosis in buffaloes is an important disease and leads to significant financial losses from condemnation of edible offal, milk, skin and energy in draught power and is also transmissible to humans.3 Various survey reported the prevalence of hydatid cyst in slaughtered buffaloes from 0.89% to 57.76% in different parts of Iran.4-8
To date, ten distinct genotypes (G1-G10) of Echinococcusgranulosus have been recognized.9,10 According to recent taxonomic revisions the E. granulosus has been ordered into 4 species namely E. granulosus sensu stricto (G1-G3), E. equines (G4), E. ortleppi (G5)and E. canadensis (G6-G10).11 The water buffalo is mainly susceptible to the G3 strain, although the sheep strain G1 and cattle strain G5 can infect this animal.12 Echinococcus granulosus strains could have important implications for host specificity, antigenicity, transmission dynamics, infection route, pathology, control, antimicrobial susceptibility, life-cycle patterns of parasite, developmental rates, fertility of developed cysts, biochemistry, infectivity in human, diagnostic reagents and vaccine development strategies.13Previousstudy mainly described the morphological and molecular characterization of E. granulosus isolates from sheep, goat, cattle, camel and human in different parts of Iran including north, center and south,14 however, there is limited molecular study dealing with the hydatid cyst from water buffaloes, in Khuzestan province, southwest Iran. This paper described a new microvariant of the G3 genotype for E. granulosus obtained from buffalo originating from southwest of Iran using comparative sequence analysis of cox1 mitochondrial gene.
Materials and Methods
Hydatid cyst was collected from liver of a 2-year-old male buffalo slaughtered in Khuzestan province, south-west Iran. Cyst contents were examined under light microscopy for the presence of protoscoleces. The collected protoscoleces were rinsed five times with sterile phosphate buffer saline (PBS) and then fixed and preserved in 70% (v/v) ethanol until DNA isolation. Following removal of the ethanol, DNA was extracted from protoscoleces using a commercial DNA extraction kit (DNesasy; Qiagen, Valencia, USA) according to the manufacturer’s protocol.
The mitochondrial DNA was amplified using specific primers for Cytochrome C oxidase 1 (cox1) JB3/JB4 (5´-TTT TTT GGG CAT CCT GAG GTT TAT -3´/5´-TAA AGA AAG AAC ATA ATG AAA ATG-3´).15 The 50 µL polymerase chain reaction (PCR) mixture contained 10 mM Tris-HCl, pH 8.4, 50 mM KCl, 1.5 mM MgCl2, 200 µM dNTPs, 20 pmol of each primer, 2 U Taq polymerase (Fermentas, Glen Burnie, USA), 8 µL of DNA extracted as template, and sterile distilled water. The PCR was performed in a thermocycler with the following program: an initial denaturation step at 95 ˚C for 5 min, 35 cycles of denaturation at 94 ˚C for 1 min, annealing at 55 ˚C for 1 min, and extension at 72 ˚C for 1 min, followed by a final extension at 72 ˚C for 5 min. The amplified products were purified with a PCR product purification kit (Bioneer, Alameda, USA) and sequenced directly using the capillary DNA analyzer (Model ABI 3730; Applied Biosystems, Foster City, USA). Nucleotide sequence analysis was undertaken using the national center for biotechnology information BLAST (Basic Local Alignment Search Tool) programs and databases.16 The different genotypes of E. granulosus, G1-G10 were used for phylogenetic analysis. Multiple sequence alignments and construction of a phylogenetic tree were made with the maximum-parsimony method using the MEGA software (Version 4.0; Biodesign Institute, Tempe, USA).17
Result
Microscopically, cyst was found to be fertile and PCR was positive by producing a cox1 fragment from DNA of E. granulosus as shown in Figure 1.

Fig. 1. Electrophoresis analysis of cox1 (493 bp). PCR amplification provided from water buffalo sample (Lane 3 and 4: duplicate samples were used for sequencing) compared with the molecular weight marker (Lane 2: 100 bp) and positive control (Lane 5). Lane 1 is negative control.
Echinococcus granulosus isolate described here was found to have complete identity with the cox1 fragment previously described by Bowles et al. for G3 reference sequence except at one nucleotide position.14 The difference was a transition mutation of adenine to guanine in position 214 (A214G) resulting in a substitution of the threonine (ACT) by alanine (GCT) according to the echinoderm mitochondria genetic code (Fig. 2). Also, phylogenetic analysis of concatenated data showed this isolate was grouped into a distinct cluster corresponding to the G1-G3 complex with the most closely relation to G3 genotype (Fig. 3).
Fig. 2. Partial chromatogram of the cox1 sequence. Dark arrow showing sense mutation of adenine to guanine in position 214.
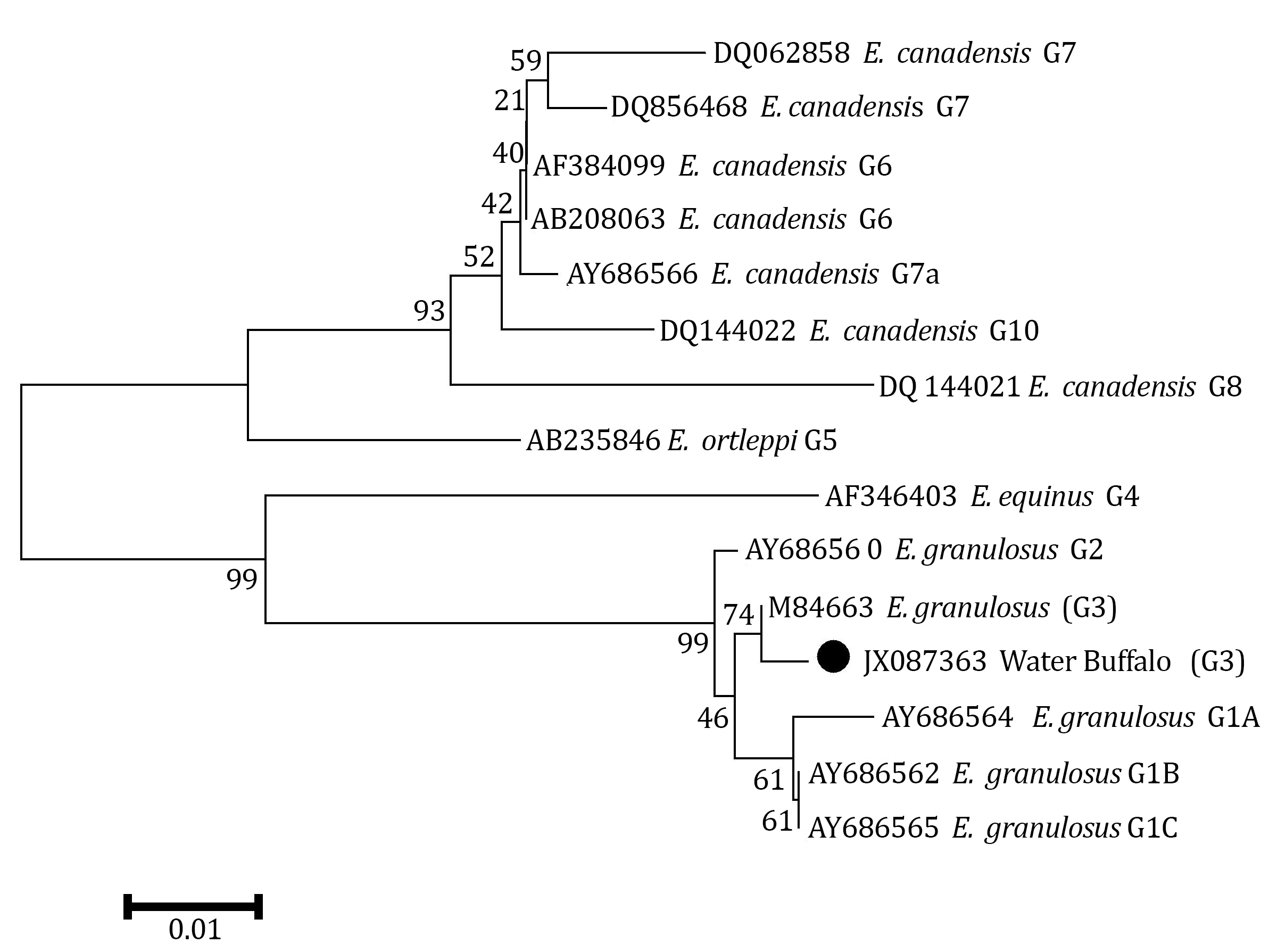
Fig. 3. A phylogenetic tree of Echinococcus granulosus genotypes inferred from the nucleotide sequences of partial cox1 and the maximum-parsimony method. Numerals indicate bootstrap values (%) from 1,000 replicates.
Discussion
According to the latest available data, there are about 460 thousand of buffaloes in Iran with approximately 138 thousand of them in the southwest region.18 Hydatidosis is an important zoonotic disease that constitutes a major public health in many countries throughout the world such as Iran.19 The application of molecular tools to the characterization of the etiological agents of echinococcosis has revealed a series of largely host-adapted species and genotypes that are maintained in distinct cycles of transmission.20 Camels and buffalo were found to have the highest prevalence of hydatid cysts. This is most likely due to the older age at which the animals are slaughtered.21 In order to develop preventive and control strategies for echinococcosis, a better knowledge of transmission cycle of E. granulosus complex is necessary. Intraspecific variants have been described for E. granulosus complex from different species of intermediate hosts in different geographical areas and several strains.20
Some investigators in various parts of the world have carried out molecular studies of the parasite. Molecular techniques have validated the genetic basis of important morphological differences that can now be used with confidence as a reliable and simple means of identifying and differentiating between strains and species of Echinococcus.22The genotypes are also important regarding the host specificity and life cycle of the E. granulosus.23,24In the current study we used a mitochondrial marker for phylogenetic studies and population differentiation because of its relatively rapid rate of evolution, importance in differentiation and discrimination of closely related geno-types (i.e. G2 and G3). It is maternally inherited and does not undergo any recombination.11,25-28 For the first time we showed the presence of the G3 genotype in water buffalo in southwest of Iran. Recently, Amin-Pour et al. reported two isolates of G3 genotypes which have 100% identity to reference G3 strain (M84663) in buffaloes generatingfrom West Azerbaijan, where this province has common border with east and southeast of Turkey.26 They could not found any G3 genotypes among buffalo samples (n = 3) obtained from southwest of Iran. Interestingly, Vural et al. have also reported G3 strain from buffaloes in east and southeast of Turkey.28 Moreover, G3 genotype was first recovered in camel in center of Iran.29 Also, this strain was found in sheep and cattle in Italy.30
According to the cox1 gene, our G3 nucleotide sequence (JX087363) differed from G3 nucleotide reference sequence (M84662), in one nucleotide at position 214 (Fig. 2). Although the sequences reported by Amin-Pour et al. and Sharbatkhori et al. for the Iranian G3 genotype are completely homologous to the reference sequence of G3 (M84663),26,29,31,32 a silent mutation (C to T at position 168) was detected in the cox1 sequences of the Iranian G3 camel isolates (HM626405) reported by Sharifiyazdi et al.27
More recently, Parsa et al. reported G3 sequences in dogs from Iran that showed 100% homology with G3 reference sequence in nad1 (AJ237633), but displayed two different cox1 profiles, each having 99.00% homology with reference G3 sequence (M84663).33 However, our study indicated that the Khuzestan strain of G3 is genetically differentiated from other G3 isolates reported so far from different geographic regions of the world. It is assumed that E. granulosus was most likely introduced into southwest Iran from Iraq through livestock transportation. Thus, additional analyses for specimens collected from neighboring countries of Iran are needed to evaluate how various G3 genotypes has been introduced and spread in Iran.
In Tunisia, M'rad et al. showed a mutation of a cytosine to a thymine in position 44 (C44T) resulting in a substitution of the alanine in position 15 by a valine according to the echinoderm mitochondrial genetic code on G3.34 In addition, Beyhan and Umur reported that six out of E. granulosus isolates from water buffalo in Turkey belonged to G1 genotypes while other samples showed variant genotypes of G1-G2-G3 complex.35 Two of them showed a thymine in position 52, and one isolate showed a single nucleotide change compared to strain G1 (C122T).
It seems that all of these mutations introduce new genotypes in the E. granulosus population and may lead to the adaptation of populations to their local environmental conditions. The investigation described here extended the information about the distribution of G3 genotype of E. granulosus in Iran and identified a new microvariant of this genotype which could be important for hydatid control and public health.
Acknowledgements
We would like to thank the authorities of School of Veterinary Medicine, Shiraz University, Shiraz, Iran for the financial support (Grant number: 87GRVT47).
| Article View | 2,160 |
| PDF Download | 1,870 |